User Codes
Astrophysics
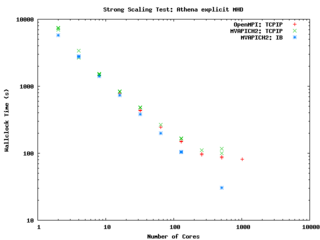
Athena (explicit, uniform grid MHD code)
Athena is a straightforward C code which doesn't use a lot of libraries so it is pretty straightforward to build and compile on new machines.
It encapsulates its compiler flags, etc in an Makeoptions.in file which is then processed by configure. I've used the following additions to Makeoptions.in on TCS and GPC:
<source lang="sh"> ifeq ($(MACHINE),scinettcs)
CC = mpcc_r LDR = mpcc_r OPT = -O5 -q64 -qarch=pwr6 -qtune=pwr6 -qcache=auto -qlargepage -qstrict MPIINC = MPILIB = CFLAGS = $(OPT) LIB = -ldl -lm
else ifeq ($(MACHINE),scinetgpc)
CC = mpicc LDR = mpicc OPT = -O3 MPIINC = MPILIB = CFLAGS = $(OPT) LIB = -lm
else ... endif endif </source> It performs quite well on the GPC, scaling extremely well even on a strong scaling test out to about 256 cores (32 nodes) on Gigabit ethernet, and performing beautifully on InfiniBand out to 512 cores (64 nodes).
-- ljdursi 19:20, 13 August 2009 (UTC)
FLASH3 (Adaptive Mesh reactive hydrodynamics; explict hydro/MHD)
FLASH encapsulates its machine-dependant information in the FLASH3/sites directory. For the GPC, you'll have to
module load intel module load openmpi module load hdf5/183-v16-openmpi
and with that, the following file (sites/scinetgpc/Makefile.h) works for me: <source lang="sh">
- Must do module load hdf5/183-v16-openmpi
HDF5_PATH = ${SCINET_HDF5_BASE} ZLIB_PATH = /usr/local
- ----------------------------------------------------------------------------
- Compiler and linker commands
- We use the f90 compiler as the linker, so some C libraries may explicitly
- need to be added into the link line.
- ----------------------------------------------------------------------------
- modules will put the right mpi in our path
FCOMP = mpif77 CCOMP = mpicc CPPCOMP = mpiCC LINK = mpif77
- ----------------------------------------------------------------------------
- Compilation flags
- Three sets of compilation/linking flags are defined: one for optimized
- code, one for testing, and one for debugging. The default is to use the
- _OPT version. Specifying -debug to setup will pick the _DEBUG version,
- these should enable bounds checking. Specifying -test is used for
- flash_test, and is set for quick code generation, and (sometimes)
- profiling. The Makefile generated by setup will assign the generic token
- (ex. FFLAGS) to the proper set of flags (ex. FFLAGS_OPT).
- ----------------------------------------------------------------------------
FFLAGS_OPT = -c -r8 -i4 -O3 -xSSE4.2 FFLAGS_DEBUG = -c -g -r8 -i4 -O0 FFLAGS_TEST = -c -r8 -i4
- if we are using HDF5, we need to specify the path to the include files
CFLAGS_HDF5 = -I${HDF5_PATH}/include
CFLAGS_OPT = -c -O3 -xSSE4.2 CFLAGS_TEST = -c -O2 CFLAGS_DEBUG = -c -g
MDEFS =
.SUFFIXES: .o .c .f .F .h .fh .F90 .f90
- ----------------------------------------------------------------------------
- Linker flags
- There is a seperate version of the linker flags for each of the _OPT,
- _DEBUG, and _TEST cases.
- ----------------------------------------------------------------------------
LFLAGS_OPT = -o LFLAGS_TEST = -o LFLAGS_DEBUG = -g -o
MACHOBJ =
MV = mv -f
AR = ar -r
RM = rm -f
CD = cd
RL = ranlib
ECHO = echo
</source>
-- ljdursi 22:11, 13 August 2009 (UTC)
Aeronautics
Chemistry
Climate Modelling
Medicine/Bio
High Energy Physics
Structural Biology
Molecular simulation of proteins, lipids, carbohydrates, and other biologically relevant molecules.
Molecular Dynamics (MD) simulation
DESMOND
GROMACS
Download and general information: http://www.gromacs.org
Search the mailing list archives: http://oldwww.gromacs.org/swish-e/search/search2.php
Compiling and Running GROMACS on Scinet (general information)
Chris Neale has compiled gromacs on GPC, with assistance from Scott Northrup, and on the power6 cluster with assistance from Ching-Hsing Yu. Users are welcome to utilize these binary executables, but only at their own peril since compiling and testing your own executable is safer and more stable.
Gromacs executables:
GPC: /scratch/cneale/exe/intel/gromacs-4.0.5/exec/bin
TCS: /scratch/cneale/exe/gromacs-4.0.4_aix/exec/bin
Below you will find, in order, scripts for the different compilations that you can follow to make your own binaries.
- Compiling serial single precision gromacs on GPC
- Compiling openmpi parallel gromacs on GPC
- Compiling serial gromacs on the power6 (submitted to the queue):
- Compiling parallel gromacs on the power6 (submitted to the queue):
- fftw single precision compilation
- Change to get mvapich2-1.4rc1 to compile gromacs
- Compiling mvapich2-1.4rc1
- Compiling gromacs on GPC using mvapich2-1.4rc1
- Submitting an IB GPC job using openmpi
- Submitting an IB GPC job using mvapich2-1.4rc1
Compiling serial single precision gromacs on GPC
<source lang="sh"> cd /scratch/cneale/exe/intel/gromacs-4.0.5 mkdir exec module purge module load intel export FFTW_LOCATION=/scratch/cneale/exe/intel/fftw-3.1.2/exec export GROMACS_LOCATION=/scratch/cneale/exe/intel/gromacs-4.0.5/exec export CPPFLAGS=-I$FFTW_LOCATION/include export LDFLAGS=-L$FFTW_LOCATION/lib ./configure --prefix=$GROMACS_LOCATION --without-motif-includes --without-motif-libraries --without-x --without-xml >output.configure 2>&1 make >output.make 2>&1 make install >output.make_install 2>&1 make distclean </source>
Compiling openmpi parallel gromacs on GPC:
<source lang="sh"> cd /scratch/cneale/exe/intel/gromacs-4.0.5 mkdir exec module purge module load openmpi intel export FFTW_LOCATION=/scratch/cneale/exe/intel/fftw-3.1.2/exec export GROMACS_LOCATION=/scratch/cneale/exe/intel/gromacs-4.0.5/exec export CPPFLAGS="-I$FFTW_LOCATION/include -I/scinet/gpc/mpi/openmpi/1.3.2-intel-v11.0-ofed/include -I/scinet/gpc/mpi/openmpi/1.3.2-intel-v11.0-ofed/lib" export LDFLAGS=-L$FFTW_LOCATION/lib /gpc/mpi/openmpi/1.3.2-intel-v11.0-ofed/lib/openmpi -I/scinet/gpc/x1/intel/Compiler/11.0/081/lib/intel64 -I/scinet/gpc/x1/intel/Compiler/11.0/081/mkl/lib/em64t/" ./configure --prefix=$GROMACS_LOCATION --without-motif-includes --without-motif-libraries --without-x --without-xml --enable-mpi --program-suffix="_openmpi" >output.configure.mpi 2>&1 make >output.make.mpi 2>&1 make install-mdrun >output.make_install.mpi 2>&1 make distclean </source>
Compiling serial gromacs on the power6 (submitted to the queue)
Note that the -O5 flag for the power6 compilation makes it take about 20 hours to compile. You can drop that if you want, but it does give you a few more percent performance.
<source lang="sh">
- ======================================================================
- Specifies the name of the shell to use for the job
- @ shell = /usr/bin/ksh
- @ job_type = serial
- @ class = verylong
- # @ node = 1
- # @ tasks_per_node = 1
- @ output = $(jobid).out
- @ error = $(jobid).err
- @ wall_clock_limit = 40:00:00
- =====================================
- this is necessary in order to avoid core dumps for batch files
- which can cause the system to be overloaded
- ulimits
- @ core_limit = 0
- =====================================
- necessary to force use of infiniband network for MPI traffic
- TURN IT OFF # @ network.MPI = csss,not_shared,US,HIGH
- necessary to force use of infiniband network for MPI traffic
- =====================================
- @ environment=COPY_ALL
- @ queue
export PATH=/usr/lpp/ppe.hpct/bin:/usr/vacpp/bin:.:/usr/bin:/etc:/usr/sbin:/usr/ucb:/usr/bin/X11:/sbin:/usr/java14/jre/bin:/usr/java14/bin:/usr/lpp/LoadL/full/bin:/usr/local/bin export F77=xlf_r export CC=xlc_r export CXX=xlc++_r export FFLAGS="-O5 -qarch=pwr6 -qtune=pwr6" export CFLAGS="-O5 -qarch=pwr6 -qtune=pwr6" export CXXFLAGS="-O5 -qarch=pwr6 -qtune=pwr6" export FFTW_LOCATION=/scratch/cneale/exe/fftw-3.1.2_aix/exec export GROMACS_LOCATION=/scratch/cneale/exe/gromacs-4.0.4_aix/exec export CPPFLAGS=-I$FFTW_LOCATION/include export LDFLAGS=-L$FFTW_LOCATION/lib cd /scratch/cneale/exe/gromacs-4.0.4_aix mkdir exec ./configure --prefix=$GROMACS_LOCATION --without-motif-includes --without-motif-libraries --without-x --without-xml >output.configure 2>&1 make >output.make 2>&1 make install >output.make_install 2>&1 make distclean </source>
Compiling parallel gromacs on the power6 (submitted to the queue)
<source lang="sh">
- ===============================================================================
- Specifies the name of the shell to use for the job
- @ shell = /usr/bin/ksh
- @ job_type = serial
- @ job_type = parallel
- @ class = verylong
- @ node = 1
- @ tasks_per_node = 1
- @ output = $(jobid).out
- @ error = $(jobid).err
- @ wall_clock_limit = 40:00:00
- =====================================
- this is necessary in order to avoid core dumps for batch files
- which can cause the system to be overloaded
- ulimits
- @ core_limit = 0
- =====================================
- necessary to force use of infiniband network for MPI traffic
- TURN IT OFF # @ network.MPI = csss,not_shared,US,HIGH
- necessary to force use of infiniband network for MPI traffic
- =====================================
- @ environment=COPY_ALL
- @ queue
export F77=xlf_r export CC=xlc_r export CXX=xlc++_r export FFLAGS="-O5 -qarch=pwr6 -qtune=pwr6" export CFLAGS="-O5 -qarch=pwr6 -qtune=pwr6" export CXXFLAGS="-O5 -qarch=pwr6 -qtune=pwr6" export FFTW_LOCATION=/scratch/cneale/exe/fftw-3.1.2_aix/exec export GROMACS_LOCATION=/scratch/cneale/exe/gromacs-4.0.4_aix/exec export CPPFLAGS=-I$FFTW_LOCATION/include export LDFLAGS=-L$FFTW_LOCATION/lib cd /scratch/cneale/exe/gromacs-4.0.4_aix echo "cn-r0-10" > ~/.rhosts echo localhost > ~/host.list for((i=2;i<=16;i++)); do
echo localhost >> ~/host.list
done export MP_HOSTFILE=~/host.list ./configure --prefix=$GROMACS_LOCATION --without-motif-includes --without-motif-libraries --without-x --without-xml --enable-mpi --disable-nice --program-suffix="_mpi" CC=mpcc_r F77=mpxlf_r > output.configure_mpi 2>&1 make mdrun > output.make_mpi 2>&1 make install-mdrun > output.make_install_mpi 2>&1 make distclean </source>
fftw single precision compilation
FFTW is required by GROMACS. This compilation must be completed before compiling GROMACS.
<source lang="sh"> mkdir exec export FFTW_LOCATION=/scratch/cneale/exe/intel/fftw-3.1.2/exec module purge module load openmpi intel ./configure --enable-float --enable-threads --prefix=${FFTW_LOCATION} make make install make distclean </source>
Change to get mvapich2-1.4rc1 to compile gromacs
This change is required to the mvapich2-1.4rc1 source code in order to compile GROMACS with it.
<source lang="sh"> src/mpid/ch3/channels/mrail/src/gen2/ibv_channel_manager.c line 503 unsigned long debug = 0; to static unsigned long debug = 0; </source>
Compiling mvapich2-1.4rc1
<source lang="sh"> cd /scratch/cneale/exe/mvapich2-1.4rc1 mkdir exec module purge module load intel ./configure --prefix=/scratch/cneale/exe/mvapich2-1.4rc1/exec CC=icc CXX=icpc F90=ifort F77=ifort >output.configure 2>&1 make >output.make 2>&1 make install >output.make_install 2>&1 make distclean </source>
Compiling gromacs on GPC using mvapich2-1.4rc1
<source lang="sh">
- !/bin/bash
cd /scratch/cneale/exe/intel/gromacs-4.0.5 mkdir exec PATH=/usr/lib64/qt-3.3/bin:/usr/kerberos/sbin:/usr/kerberos/bin:/usr/local/sbin:/usr/local/bin:/sbin:/bin:/usr/sbin:/usr/bin:/opt/xcat/bin:/opt/xcat/sbin:/root/bin:/opt/torque/bin:/opt/xcat/bin:/opt/xcat/sbin:/usr/lpp/mmfs/bin:/scratch/cneale/exe/mvapich2-1.4rc1/exec/bin/:/scinet/gpc/x1/intel/Compiler/11.0/081/bin/intel64 LD_LIBRARY_PATH=/scratch/cneale/exe/mvapich2-1.4rc1/exec/lib/:/scinet/gpc/x1/intel/Compiler/11.0/081/lib/intel64:/scinet/gpc/x1/intel/Compiler/11.0/081/mkl/lib/em64t/ export FFTW_LOCATION=/scratch/cneale/exe/intel/fftw-3.1.2/exec export GROMACS_LOCATION=/scratch/cneale/exe/intel/gromacs-4.0.5/exec export CPPFLAGS="-I$FFTW_LOCATION/include -I/scratch/cneale/exe/mvapich2-1.4rc1/exec/include -I/scratch/cneale/exe/mvapich2-1.4rc1/exec/lib" export LDFLAGS=-L$FFTW_LOCATION/lib ./configure --prefix=$GROMACS_LOCATION --without-motif-includes --without-motif-libraries --without-x --without-xml --enable-mpi --program-suffix="_mvapich2" >output.configure.mpi.mvapich2 2>&1 make >output.make.mpi.mvapich2 2>&1 make install-mdrun >output.make_install.mpi.mvapich2 2>&1 make distclean </source>
Submitting an IB GPC job using openmpi
<source lang="sh">
- !/bin/bash
- PBS -l nodes=10:ib:ppn=8,walltime=40:00:00,os=centos53computeA
- PBS -N 1
if [ "$PBS_ENVIRONMENT" != "PBS_INTERACTIVE" ]; then
if [ -n "$PBS_O_WORKDIR" ]; then cd $PBS_O_WORKDIR fi
fi /scinet/gpc/mpi/openmpi/1.3.2-intel-v11.0-ofed/bin/mpirun -np $(wc -l $PBS_NODEFILE | gawk '{print $1}') -machinefile $PBS_NODEFILE /scratch/cneale/exe/intel/gromacs-4.0.5/exec/bin/mdrun_openmpi -deffnm pagp -nosum -dlb yes -npme 24 -cpt 120
- To submit type: qsub this.sh
</source>
Submitting an IB GPC job using mvapich2-1.4rc1
Note that mvapich2-1.4rc1 is not configured to fall back to ethernet so this will not work on the non-IB nodes, even for 8 cores.
<source lang="sh">
- !/bin/bash
- PBS -l nodes=4:ib:ppn=8,walltime=30:00:00,os=centos53computeA
- PBS -N 1
if [ "$PBS_ENVIRONMENT" != "PBS_INTERACTIVE" ]; then
if [ -n "$PBS_O_WORKDIR" ]; then cd $PBS_O_WORKDIR fi
fi module purge module load mvapich2 intel /scratch/cneale/exe/mvapich2-1.4rc1/bin/mpirun_rsh -np $(wc -l $PBS_NODEFILE | gawk '{print $1}') -hostfile $PBS_NODEFILE /scratch/cneale/exe/intel/gromacs-4.0.5/exec/bin/mdrun_mvapich2 -deffnm pagp -nosum -dlb yes -npme 12 -cpt 120
- To submit type: qsub this.sh
</source>
Things still left to do for GROMACS
Intel has it's own fast fourier transform library, which we expect to yield improved performance over fftw. We have not yet attempted such a compilation.
GROMACS benchmarks on Scinet
This is a rudimentary list of scaling information.
I have a 50K atom system running performance on GPC right now. On 56 cores connected with IB I am getting 55 ns/day. I set up 50 such simulations, each with 2 proteins in a bilayer and I'm getting a total of 5.5 us per day. I am using gromacs 4.0.5 and a 5 fs timestep by fixing the bond lengths and all angles involving hydrogen.
I can get about 12 ns/day on 8 cores of the non-IB part of GPC -- also excellent.
As for larger systems, My speedup over saw.sharcnet.ca for a 1e6 atom system is only 1.2x running on 128 cores in single precision. Although saw.sharcnet.ca is composed of xeons, they are running at 2.83 GHz (https://www.sharcnet.ca/my/systems/show/41), which is a faster clock speed than the Scinet 2.5 GHz for Intel's next-generation X86-CPU architecture. While GROMACS is generally not excellent for scaling up to or beyond 128 cores (even for large systems), our benchmarking of this system on saw.sharcnet.ca indicated that it was running at about 65% efficiency. Benchmarking was also done on Scinet for this system, but was not recorded as we were mostly tinkering with the -npme option to mdrun in an attempt to optimize it. My recollection, though, is that the scaling was similar on scinet.
-- cneale 18 August 2009